Mapping the climatic gradients driving landscape genomic variation
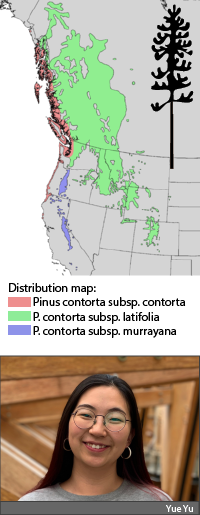
Climate is a major environmental factor driving genetic variation of forest trees across the landscape. Associating the genetic variation in adaptive traits with climatic variables has been increasingly important for developing forest resource management strategies to adapt to climate change, such as climate-based seed transfer guidelines. However, such studies require long-term common garden experiments that are expensive to establish and maintain, also time-consuming. Rapid advances in genomics may provide an alternative solution despite that associating genomic variation with adaptive traits remains a challenge. In my research project for my Master’s thesis (under Dr. Wang‘s supervision), the key objective is to explore the climate variables that drive genetic variation in lodgepole pine over the landscape. Using the lodgepole pine SNP data generated from the AdapTree project completed in Aitken’s lab and the climate data from ClimateNA. I am trying to model the underlying relationship between genomic variation and climate variables using machine-learning approaches including GradientForest. I will determine the relationship between genetic variation (SNPs loci) and important predictors (climate variables) identified through the modeling process. Variations in SNPs are aggregated by GradientForest to identify the composition and turnovers along each of these environmental gradients. This project would contribute to a better understanding of climate adaptation of forest tree species and provide information for developing forest adaptive strategies.
Funding: China Scholarship Council, NSERC Discovery Grant (RGPIN-2018-04643)
